Reading Remote Data Files
Sometimes data arrives as a series of individual files each of which is organized in the same way—which is to say, each of which has the same variables, features, or columns. Imagine a series of tables reporting mandated information about every school in the state, or a hundred spreadsheets each with information about a different country, or thirty seven CSVs each with the same columns of information about representatives in each U.S. Congressional Session since 1945. Seeing as these data were not provided to us as a single big table we generally want to make it into one. If the files are CSVs on our local computer, R has some nice functions that allow us to iterate over a vector of filenames and produce a tidy table of data.
Case 1: Lots of local CSV files
For example, our Congressional project might have a data folder with a subfolder called congress. We can get a listing of the CSV files inside it like this:
|
|
We can feed that vector to read_csv() and it will be quite happy. It reads each file and binds them all together by row, in a big stack. (Again, remember we know ex ante that the files all have the same column structure. We’ll get errors or warnings if this isn’t true.)
|
|
You can see how path is created as an id column, to help us keep track of which file each row of data came from. We create the congress column after we read in the data by extracting the congressional session from the filename with a regular expression. But you can see how nice it is to have this facility to read data in like this.
Case 2: Lots of local Excel files
What if the data is in a file format whose read-in function doesn’t know this trick about accepting a vector of file paths? In that case we can do what read_csv() is doing behind the scenes and map the vector of file names to the read-in function, and explicitly bind the results together. The default map() function binds by making a list of whatever you did. So for example, if our Congressional data were all Excel files in .xlsx format, and they all had the same structure with a header row of column names, we could write this:
|
|
And we would get (almost) the same result. Here we start with the vector of filenames and pass it down a pipe to set_names(), which usefully labels the elements of the filename vector with their values (in this case, the file paths). Then we pass it along to map(), which goes ahead and maps, or applies, the read_xlsx() function to each element of the filenames vector—i.e. to each file. The \(x) notation there is a the shorthand for an anonymous function. Each element of the filenames vector becomes the x that is read in turn. Mapping a function like this is just a kind of iteration where you don’t have to explicitly write a loop. This makes it easier to cleanly compose sequences or chains of functions without having to create a bunch of placeholder objects, declare counters, and so on.
Case 3: Lots of remote CSV or Excel files in a bare directory
Now, what if the files we want are stored remotely on a server? These days there’s often an API for such things. But quite often, even now, you may find yourself dealing (as I did yesterday) with a bare directory of files that looks like this:
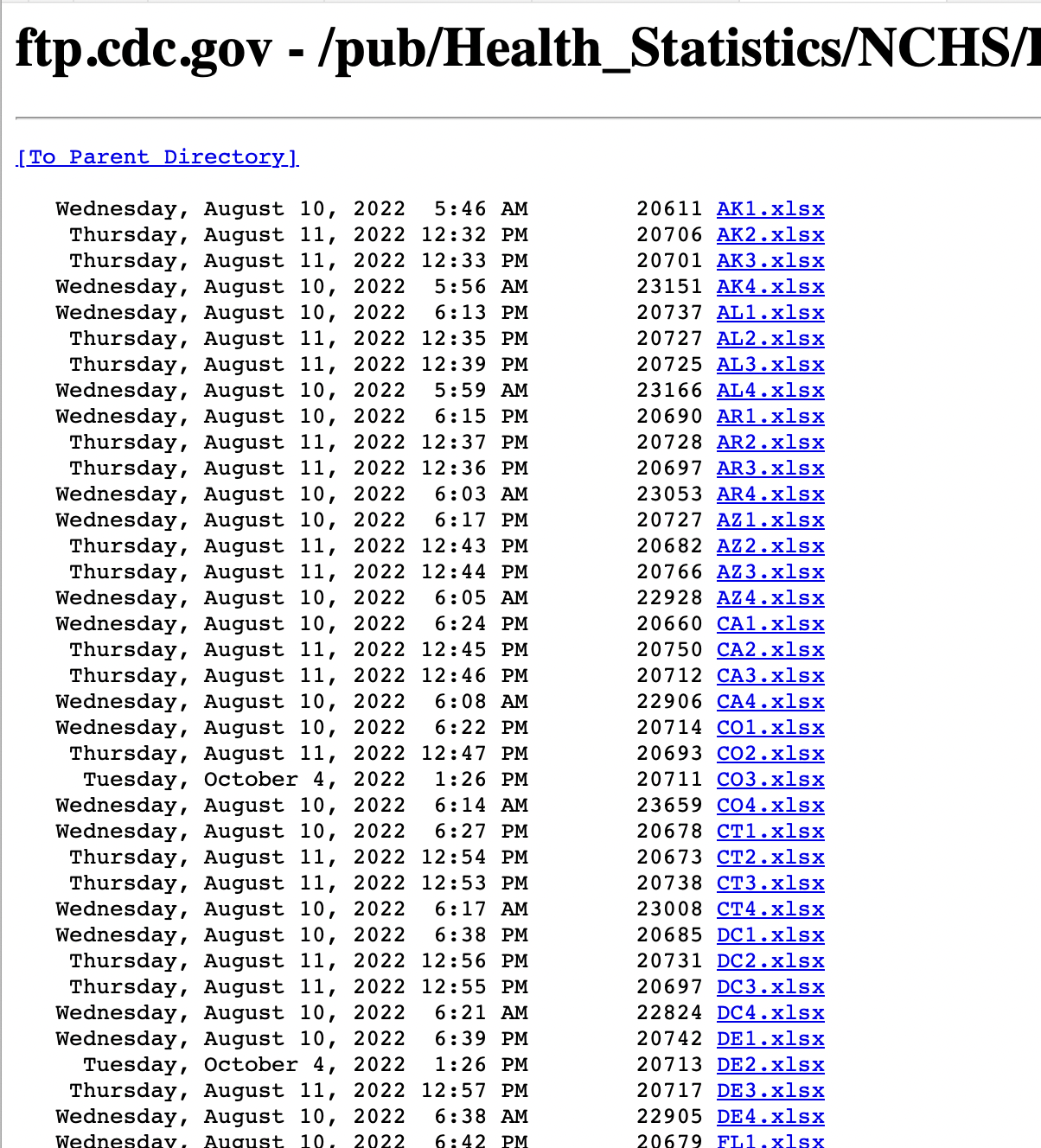
A listing of state-level life-tables from the National Center for Health Statistics
This is a directory of state-by-state life tables associated with a CDC report. Again, we want them (or, as we’ll see, some of them) as a single table.
Now, if these were provided as CSVs our task would be a little easier because in addition to being able to deal with a vector of filenames at once, read_csv(), and indeed all the read-in functions in readr in general, will happily read URLs as well as local file paths. However, the read_xlsx() function in the readxl package can’t do this yet. It only wants file paths. A second issue is that the Excel files themselves are not entirely tidy. At the top they look like this:

The top of a state-level life table Excel file.
Those first two rows are a mild violation of one of the rules of thumb for entering data in spreadsheets, helpfully outlined by Karl Broman and Kara Woo. The first row is metadata; the second is a more verbose description of the standard lifetable headers in the third row. Except for Age, which is not labeled in the third row. That big box labeled “Age (years)” is actually an super-sized second row. That means the first element of row three, our actual column headers, is blank! This is annoying. Fortunately these are easily dealt with, as we can just tell our read function to skip those two lines. There’s also a Source row at the bottom (not shown here) that we’ll have to strip out.
But the first order of business is getting a vector of the actual file URLs to download. You could just copy and paste the listing, like an animal, but we are not going to do that. Instead, we’ll look at two ways to get the file listing programmatically. First, we’ll take advantage of the old-school ftp-style directory listing to get the file names. We’ll do this using R’s implementation of curl.
|
|
We open a connection to the remote folder and use ftp_use_epsv and dirlistonly flags to restrict what we get back. Then we read the lines recieved from the FTP server into the files object. This gives us the bare file names of everything in this remote folder.
The spreadsheets are named according to a scheme with a two-letter state abbreviation followed by a number. The number signifies the kind of life-table it is. The files ending in 1 have the life-table for the population as a whole, which is what we are interested in.
Literally the day after I wrote this example, the CDC turned off FTP access to these pages. Now they are HTTPS only. So the code above no-longer works. However, alternatively and a little more straightforwardly we can use rvest to get the filenames, extract all the link elements, and get the text from inside of them. The rvest package handles the business of opening and closing web connections for us. It also provides handy functions to get and extract the text of pages based on there position in a CSS selection hierarchy (which isn’t relevant here) or based on particular HTML elements. First we get the page, then we grab all the link elements in it, and then convert their content to a character vector:
|
|
In this case we’d need to clean up the resulting vector (to remove navigation links to the parent directory and so on) but would end up in the same place.
Now that we have a vector of the file names (but just the file names at this point) we can do a bit of prep. As you can see we use set_names() again. It’s very handy. This time we make each element’s name attribute be the state abbreviation rather than the full URL.
|
|
The label makes it much easier to create a state id column, because after we map(), list_rbind() can use the label as its index counter.
The last step is to get read_xlsx() to get all the remote files, which it does not have the capacity to do directly. It won’t even accept a single URL, it only wants file paths. So we will have to write a one-off function that gets the file and puts it in a temporary location that read_xlsx() can see.
|
|
The first line inside the function uses httr to GET the file, and immediately save it locally using write_disk(), taking care to specify that the temporary file we save should have an .xlsx extension. (Otherwise read_xlsx() will complain.) The second line actually reads in the file that’s been downloaded. We take the opportunity to suppress chatter about the name repair that has to happen on that blank first column header in the third row, rename that location age, and strip the trailing line about the source of the data that I mentioned above.
This function reads one given URL. Now we just need to map a vector of URLs to it and bind the results by row:
|
|
The map() function makes a list of all the data frames and list_rbind() binds them. As I said, a nice thing is that will use the name attribute of fnames to create its id column, which we can therefore name state, because the name of each URL element is the abbreviation for the state it is providing data about.
And we’re done:
|
|
A few thousand rows of data programmatically extracted from fifty spreadsheets, now ready for a bit more cleaning and any amount of confusion about what it is that life-tables actually show.